|
12/30/2023 0 Comments Github opendriveThe process is simple, just execute this next command on any Colab notebook and follow the link that it will display in the output. Of course, you could choose to clone to that folder form an actual computer, with access to the terminal, and later on, go on to open the files on Colab (or anywhere else), but as I mentioned, I found myself without a computer (although, what’s a computer right?) and I had to figure out how to clone my repos using my iPad.įor this, you will have to start by mounting Google Drive into Google Colab, which already has git installed so at least that is covered. However, if you clone a GitHub repo to your Drive folder, you can access it anytime. 
Accessing Google Drive from Google Colabīecause by default the directories that you can access from Colab are not the ones on your Drive, it would make it very hard (if at all possible) to access those files later. But let me explain how I solved both issues and am now able to clone any repo to my Google Drive, edit my ipynb files from Colab and push everything back to GitHub even from my iPad Pro. And look, I’m sure that you could solve the credentials problem in some other way, but I was in a hurry, and so the solution may not have been the best or more secure. And also, if I tried to push back to the remote, that failed because Colab never prompted me to enter my GitHub credentials. I had managed to clone my repo, as you would, to the content folder that is enabled for your Google Colab notebook, I could see my ipynb files listed right there, and yet I was unable to open them. 
Also, when it comes time to push everything back to the remote, it simply will fail (no way to enter the credentials). 
This means that if you were to directly clone a GitHub repo, it will end up in some directory that you won’t easily find from outside Colab (if at all). The reason why cloning a GitHub repository with Google Colab is not as straight forward as one may think is twofold:īy default, you don’t have an easily accessible File System
0 Comments
12/30/2023 0 Comments Beacon designer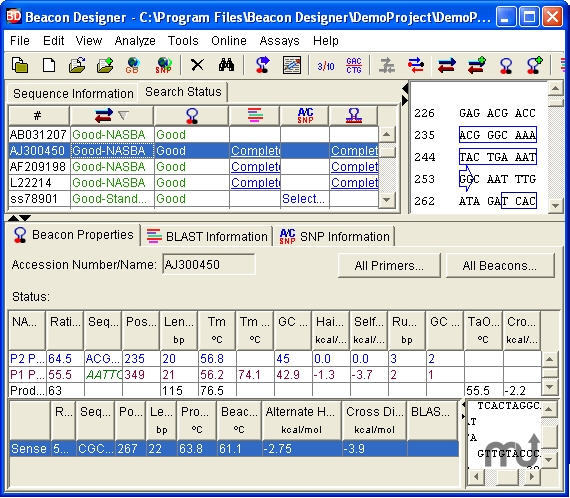
Polymerase Chain Assembly (PCA) - created to automate the design oligonucleotide sets for long sequence assembly by PCR. translates nucleotide (DNA/RNA) sequences to the corresponding peptide sequence in all six frames for standard and degenerate DNA and modifications (inosine, uridine).tool for searching for similar sequences (or primers).tool for restriction I-II-III types enzymes analysis, find or create restriction enzyme recognition sites for coding sequences.tool for identifying simple sequence repeat (SSR) loci by analysing the low complexity regions of input sequences.tool calculate Tm for primer dimers with mismatches for pure and mixed bases using averaged nearest neighbour thermodynamic parameters and for modifications (inosine, uridine, or LNA).primers (probes) are analyzed for all primer secondary structures including the alternative hydrogen bonding to Watson-Crick base pairing such as G-quadruplexes or wobble base pairs (like G-G, G-T, G-A), self-dimers and cross-dimers in primer pairs.analyzes different features of multiple primers simultaneously, the melting temperature, GC content, sequence linguistic complexity, primer PCR efficiency and molecular weight, the extinction coefficient, the optical density (OD).comprehensive primer test, the melting temperature calculation for standard and degenerate oligonucleotides including LNA and other modifications, primer's PCR efficiency and linguistic complexity, and dilution and resuspension calculator.in silico (virtual) PCR for primers and probes searching or in silico PCR against genome(s) (or a list of chromosome) - prediction of probable PCR products and search of potential mismatching location of the specified primers or probes.java application is based on FastPCR software for Windows and provides comprehensive and professional facilities for designing primers for most PCR applications and their combinations: standard, multiplex, long distance, inverse, real-time, Xtreme Chain Reaction (XCR®), group-specific (universal primers for phylogenetically related DNA sequences) or unique (specific primers for each from phylogenetically related DNA sequences), bisulphite modification assays, polymerase extension PCR multi-fragments assembly cloning (OE-PCR), Loop-mediated Isothermal Amplification (LAMP) microarray design Allele-Specific PCR (AS-PCR) - KASP™ (Kompetitive Allele Specific PCR) or PACE™ (PCR Allele Competitive Extension) based genotyping assay design for biallelic discrimination of single nucleotide polymorphisms (SNPs) and insertions and deletions (Indels) at specific loci. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
